4peaks downlad
- #4PEAKS DOWNLAD HOW TO#
- #4PEAKS DOWNLAD MANUAL#
- #4PEAKS DOWNLAD FULL#
- #4PEAKS DOWNLAD SOFTWARE#
- #4PEAKS DOWNLAD DOWNLOAD#
The trimmed sequence can be obtained here as FASTA, EzEditor2 file. I trimmed at the absolute position of 1,213 which left 519 bases. In this example, the ab1 contains 1,539 bases (very long for a single Sanger reaction). 400 bp is sufficient to get an accurate identification. Find the position where you want to trim the right-side (the rest of sequence). Compare your sequence with the hit reference sequence and also consult the chromatogram. This will move the cursor to the next position where nucleotide does not match to the hit reference sequence. A tip is to use the shortcut Ctrl+U key (Compare UP function).

Right-click to activate the popup menu then select “Delete”->”Before cursor”. Move the cursor to the position where you want to trim. You need to trim this erroneous region in the EzEditor2 tool. The corresponding region in the chromatogram also shows low quality sequencing reaction (box (b) of the blow). The sequence of the best hit ( Ruminococcus faecis) is located at the right upper row of your sequence).Īlignment your sequence against the hit sequence ( Ruminococcus faecis) by clicking (c).Īfter the automated pairwise alignment, you will see the erroneous region that does not match to other reference sequences (box (a) of the above). Check your sequence (from ab1 file) is at the bottom (b). Move the cursor (a) to the bottom of the screen. Your sequence is located at the bottom of the alignment.
#4PEAKS DOWNLAD SOFTWARE#
Start EzEditor2 software and open the file that you have just downloaded. This file contains your sequences and reference sequences that show the highest sequence similarities.
#4PEAKS DOWNLAD DOWNLOAD#
Click (c) to view the detailed result of “Identify”.Ĭlick (a) to download a data file for EzEditor2 tool. We will start the trimming process using a sequence alignment tool called EzEditor2. As you will see the chromatogram shown above, your sequence contains errors at the front end (as well as backend). This is cutoff is applicable for high-quality sequence only. The generally accepted 16S similarity cutoff is 98.7%. In this case, sequence similarity is 95.35% and you may be excited to discover a potentially new species. The hit species (a) is displayed with pairwise sequence similarity (b). Go to and paste your sequence as new query. Then, search through the EzBioCloud’s Identify. This file contains the result of the fairly good sequencing reaction. This contains a 16S sequence of a human fecal bacterium. If you used the sequence data without trimming the ends, your sequence will have a substantial amount of errors. Because of the sequencing chemistry, both ends of sequence contain substantial errors.
#4PEAKS DOWNLAD MANUAL#
Quality and manual inspection of the raw data is possible with the chromatogram.
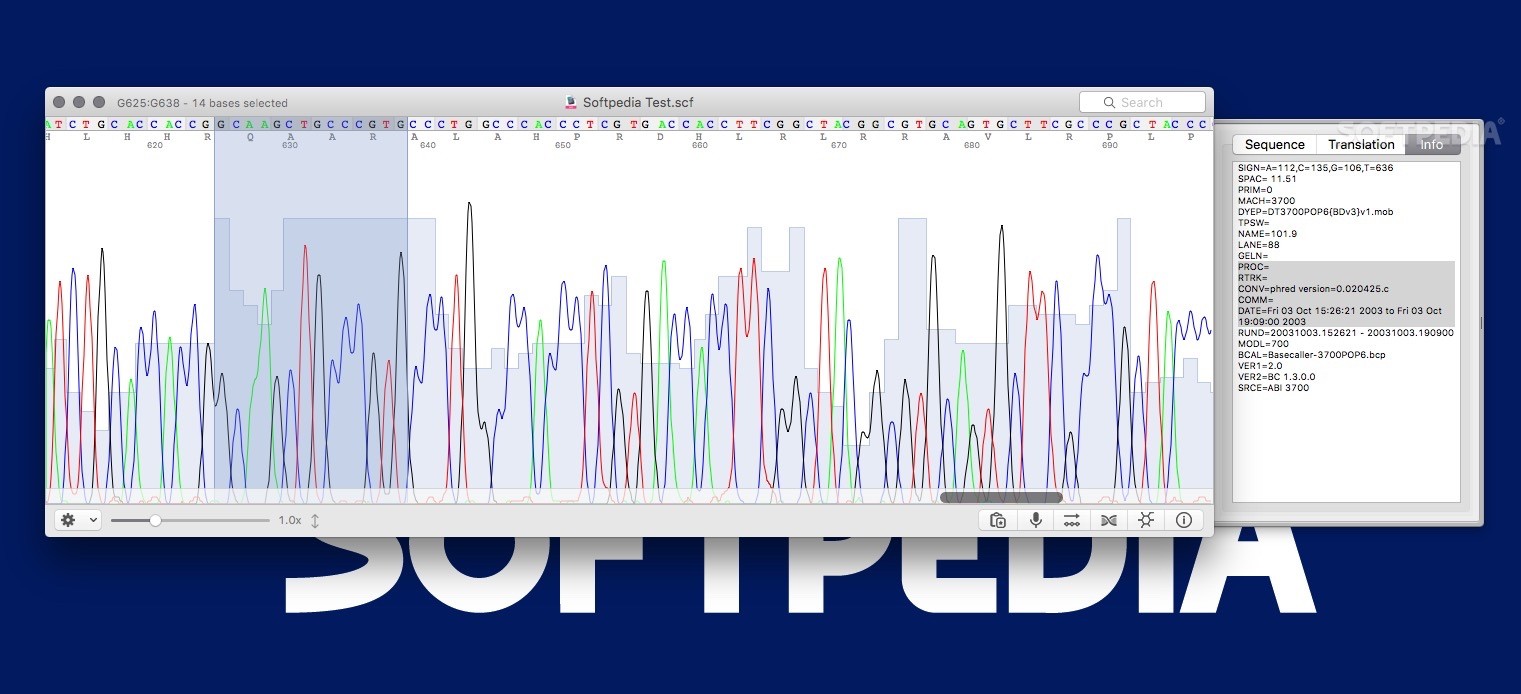
An ab1 file contains the chromatogram information from ABI sequencers ( Learn more).
#4PEAKS DOWNLAD HOW TO#
She will be looking for sponsors to help her achieve her target of R20 000 in tackling and completing this challenge for a good cause.This document explains how to use EzBioCloud’s “Identity” service using single Sanger sequencing data. Mackenzie has always loved Penguins and seabirds and has a passion to raise awareness around the plight of animals, and has set this challenge up to create support for SANNCOB and it's conservation efforts.
She will start at Suikerbossie in Hout Bay and finish at Constantia Nek targeting sub 13hrs over the 30km distance and 1500m elevation. The peaks she will summit, in order, will be Judas Peak, Grootkop, Maclears Beacon and Klaasenkop.
#4PEAKS DOWNLAD FULL#
In the March school holiday, Mackenzie will complete a 30km mountain challenge, summiting 4 peaks on the top of Table Mountain, covering the full length and width of the mountain top. After completing many mountain challenges in 2021 (including a multi-day 13 Peaks and 12hr 51min Table Mountain challenge), Mackenzie has set mountain challenges for 2022 with the goal of making a difference. Mackenzie Knott is an 8yr old student from Rallim Preparatory and shares a great passion for the outdoors, as well as a good challenge.
